According to the latest estimates a human cell has around 21,000 genes and being a multicellular organism all the cells have a copy of these genes. Presence of nucleus and complexity of eukaryotic organism demands a well controlled gene regulation in eukaryotic cell. Tissue specific gene expression is essential as they are multicellular organisms in which different cells perform different functions.
1.1 Expression of Different Genes
Different Kinds of genes in a multicellular organisms
1.2 Mechanism of Gene Regulation in Prokaryotes and Eukaryotes
The mechanism of regulation of genes is different in prokaryotes and eukaryotes. Eukaryotes have a well defined nucleus which is surrounded by a nuclear membrane. This nuclear membrane prevents the simultaneous transcription and translation which takes place in prokaryotes. Hence
In Eukaryotes the ultimate synthesis of proteins is regulated right from the chromatin stage. A number of intermediate steps lying between transcription and translation serve as various check points at which the gene expression can be controlled.
1.3.1 Chromatin structure
The folding of DNA is not uniform throughout. In some regions the folding is highly compact and intricate giving rise to heterochromatic regions and the others are called euchromatin regions.
Mechanisms which affect the chromatin structure and hence the expression of gene are:
A. Histone modifications - Please refer section on histone modifications for more details.
B. Methylation of DNA
The cytidine residues which exist as a dinucleotide, CG (written as CpG) often serve as target sites for methylation with the DNA. The CpG dinucleotides are found 10-20 times more in the promoter region than the rest of the genome. The region which has more methylated cytidine residues is transcriptionally inactive.
1.3.2 Regulation of Transcription
It is a major check point for controlling the expression of gene in both eukaryotes and prokaryotes. The differences in the mechanisms by which the transcription of gene is controlled in prokaryotes and eukaryotes are listed below:
Prokaryotes
Eukaryotes
Role of Promoters
Prokaryotes
There are two promoter elements or DNA sequences which are 35 and 10 base pairs in length and seated upstream to the transcriptional initiation sites.
The consensus sequence present at
Promoters in eukaryotes are of two types:
Enhancers can be located upstream, downstream or within the gene that is transcribed. The binding of these enhancers with enhancer binding proteins (transcription factors) increases the rate of transcription of that gene to a greater extent. Promoters are capable of initiating lower levels of transcription. Enhancers are responsible for the cell or tissue specific transcription. Each enhancer has its own transcription factor that it binds to.
1.1 Expression of Different Genes
Different Kinds of genes in a multicellular organisms
- House keeping genes - Few very essential genes known as house keeping genes are expressed in all the cells all the time.
- Genes required during cellular differentiation - Few other genes are required during differentiation helping a cell to perform certain specific functions like a skin cell, liver cell etc. Prokaryotic organisms are simple and unicellular but multicellular organisms are made up of a number of cells which perform different kinds of function. The regulation of genes is very essential for cell differentiation.
- Genes which get triggered as a response to some external factors - Genes which help in the production of hormones.
- Genes which get triggered during apoptosis- Genes which control the cell death.
1.2 Mechanism of Gene Regulation in Prokaryotes and Eukaryotes
The mechanism of regulation of genes is different in prokaryotes and eukaryotes. Eukaryotes have a well defined nucleus which is surrounded by a nuclear membrane. This nuclear membrane prevents the simultaneous transcription and translation which takes place in prokaryotes. Hence
- In prokaryotes the primary control point is the process of transcription initiation
- In eukaryotes expression of gene into proteins can be controlled at various locations.
In Eukaryotes the ultimate synthesis of proteins is regulated right from the chromatin stage. A number of intermediate steps lying between transcription and translation serve as various check points at which the gene expression can be controlled.
1.3.1 Chromatin structure
The folding of DNA is not uniform throughout. In some regions the folding is highly compact and intricate giving rise to heterochromatic regions and the others are called euchromatin regions.
Mechanisms which affect the chromatin structure and hence the expression of gene are:
A. Histone modifications - Please refer section on histone modifications for more details.
B. Methylation of DNA
The cytidine residues which exist as a dinucleotide, CG (written as CpG) often serve as target sites for methylation with the DNA. The CpG dinucleotides are found 10-20 times more in the promoter region than the rest of the genome. The region which has more methylated cytidine residues is transcriptionally inactive.
1.3.2 Regulation of Transcription
It is a major check point for controlling the expression of gene in both eukaryotes and prokaryotes. The differences in the mechanisms by which the transcription of gene is controlled in prokaryotes and eukaryotes are listed below:
Prokaryotes
- The linked genes are organized into clusters known as operons which are under the control of a single promoter.
- These genes are primarily regulated by repressors.
- A promoter sequences which controls an operon lies upstream of the operon. Accessory or the regulatory proteins control the recognition of the transcriptional initiation sites by RNA polymerases.
- A single operon gets transcribed into a polycistronic mRNA which can be translated into multiple proteins
Eukaryotes
- Eukaryotic genes are not organized into operons and each of these genes requires its own promoter.
- Regulation by repressors is very occasional and the primary role of regulation is played by the transcriptional activators known as transcription factors.
- Those genes which code for a protein have a basic structure consisting of:
- Exons – gene sequences which encode for a polypeptide
- Introns - these sequences will get removed from the mRNA before it gets translated.
- A transcription cite
- Promoter sequences.
- Monocistronic mRNAs which can produce a single polypeptide are produced
Role of Promoters
Prokaryotes
There are two promoter elements or DNA sequences which are 35 and 10 base pairs in length and seated upstream to the transcriptional initiation sites.
The consensus sequence present at
- -35 position is TTGACA
- -10 position is TATAAT. This is also termed as Pribnow-box.
Promoters in eukaryotes are of two types:
- Basal promoter or core promoter – These promoters reside within 40bp upstream of the start site. These promoters are seen in all protein coding genes. Examples are CCAAT-boxes and TATA-boxes
- TATA box
- The consensus sequence for TATA box is TATAT/AAT/A
- It resides 20 to 30 bases upstream of the transcriptional start site
- This is similar in sequence to the prokaryotic Pribnow-box
- Proteins like TFIIA, B, C interact with this TATA box
- CCAAT-box
- The consensus sequence for this is GGT/CCAATCT
- It resides 50 to 130 bases upstream of the transcriptional start site
- Protein named as C/EBP (CCAAT-box/Enhancer Binding Protein) binds this box
- Upstream promoters _ these promoters may lie up to 200bp upstream of the transcriptional initiation site. The structure of this promoter and the associated binding factors keeps varying from gene to gene.
Enhancers can be located upstream, downstream or within the gene that is transcribed. The binding of these enhancers with enhancer binding proteins (transcription factors) increases the rate of transcription of that gene to a greater extent. Promoters are capable of initiating lower levels of transcription. Enhancers are responsible for the cell or tissue specific transcription. Each enhancer has its own transcription factor that it binds to.
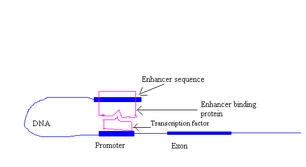
Action of an Enhancer
Enhancer binding proteins have two binding sites
How is the gene transcription controlled at this point?
The unique combination of the promoter sites, transcription factors and enhancers chosen ultimately decides which gene gets switched on and which one gets switched off.
1.3.3 Regulation of RNA Processing
After transcription, RNA is synthesized and it undergoes RNA processing which involves
Depending on the final combination of exons, achieved after splicing we can determine the nature of the protein. The function of proteins varies in cell one and two.
Enhancer binding proteins have two binding sites
- One site binds DNA
- Another site binds the transcription factors that are bound to the promoter. This draws the DNA into a loop and stabilizes it by the force of cohesion.
How is the gene transcription controlled at this point?
The unique combination of the promoter sites, transcription factors and enhancers chosen ultimately decides which gene gets switched on and which one gets switched off.
1.3.3 Regulation of RNA Processing
After transcription, RNA is synthesized and it undergoes RNA processing which involves
- Addition of 5' cap
- Addition of a 3' poly (A) tail
- Removal of introns
- Chance1 – If translation of RNA is destined then only it will get transported out of the nucleus other wise not.
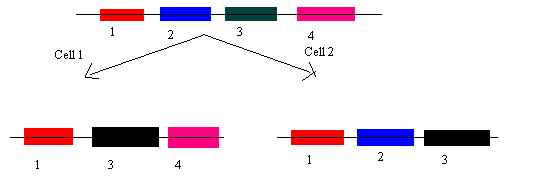
- Chance 2- Exon shuffling
Depending on the final combination of exons, achieved after splicing we can determine the nature of the protein. The function of proteins varies in cell one and two.
Exon shuffling
1.3.4 Regulation of RNA Transport
Only some RNAs function within the nucleus whereas all other RNAs which are meant for protein synthesis have to be transported from the nucleus to the cytoplasm via nuclear pores.
1.3.5 Regulation of RNA Longevity
The life spans of different mRNAs from different genes are different. The information of the lifespan of mRNA is found in the 3' UTR. The sequence AUUUA, when found in the 3' UTR, is a signal for early degradation (and therefore short lifetime). Greater is the number of times the sequence is present, the shorter the lifespan of the mRNA.
Example: The RNAs which have a shorter lifespan produce lesser number of polypeptides in comparison to those which have a longer lifespan. Hence to regulate the number of polypeptides produced a cell tries to regulate the longevity of the mRNA. Prokaryotic mRNAs, have half-lives in the range of 1 to 5 minutes.
1.3.6 Regulation of Translation
A. Translational initiation
The expression of a gene product also depends on the ability of the ribosome to recognize the correct AUG codon out of the multiple methionine codons present in the mRNA.
B. Control of translational process
A large number of RNAs are produced in a cell but they are stopped from getting translated by various mechanisms which are less understood. For example in many animals large amounts of mRNAs are produced by the eggs but all of them do not get translated until the egg is fertilized.
Every mRNA consists of three parts - 5' untranslated region (5'UTR), protein-coding region or open reading frame (ORF) and 3' untranslated region (3'UTR).
1.3.7 Post Translational Control Points
A. Post translational modifications
A polypeptide ultimately attains a functional state when it attains a three dimensional state and undergoes certain modifications like glycosylation, acetylation, fatty acylation, disulfide bond formations, etc. Enzymes known as chaperons assist the newly formed protein to attain the three dimensional structure which is needed for the proper functioning.
B. Protein transport
Many proteins need to get transported to their site of action. Due to the complexity of the cell there are various targeting processes within the eukaryotes which ensure that the protein arrives at the correct location. Some proteins like digestive enzymes or hormones even get exported out of the cell.
C. Protein stability
The lifespan of a protein depends on the specific amino acid sequence present within them.
1.3.4 Regulation of RNA Transport
Only some RNAs function within the nucleus whereas all other RNAs which are meant for protein synthesis have to be transported from the nucleus to the cytoplasm via nuclear pores.
1.3.5 Regulation of RNA Longevity
The life spans of different mRNAs from different genes are different. The information of the lifespan of mRNA is found in the 3' UTR. The sequence AUUUA, when found in the 3' UTR, is a signal for early degradation (and therefore short lifetime). Greater is the number of times the sequence is present, the shorter the lifespan of the mRNA.
Example: The RNAs which have a shorter lifespan produce lesser number of polypeptides in comparison to those which have a longer lifespan. Hence to regulate the number of polypeptides produced a cell tries to regulate the longevity of the mRNA. Prokaryotic mRNAs, have half-lives in the range of 1 to 5 minutes.
1.3.6 Regulation of Translation
A. Translational initiation
The expression of a gene product also depends on the ability of the ribosome to recognize the correct AUG codon out of the multiple methionine codons present in the mRNA.
B. Control of translational process
A large number of RNAs are produced in a cell but they are stopped from getting translated by various mechanisms which are less understood. For example in many animals large amounts of mRNAs are produced by the eggs but all of them do not get translated until the egg is fertilized.
Every mRNA consists of three parts - 5' untranslated region (5'UTR), protein-coding region or open reading frame (ORF) and 3' untranslated region (3'UTR).
1.3.7 Post Translational Control Points
A. Post translational modifications
A polypeptide ultimately attains a functional state when it attains a three dimensional state and undergoes certain modifications like glycosylation, acetylation, fatty acylation, disulfide bond formations, etc. Enzymes known as chaperons assist the newly formed protein to attain the three dimensional structure which is needed for the proper functioning.
B. Protein transport
Many proteins need to get transported to their site of action. Due to the complexity of the cell there are various targeting processes within the eukaryotes which ensure that the protein arrives at the correct location. Some proteins like digestive enzymes or hormones even get exported out of the cell.
C. Protein stability
The lifespan of a protein depends on the specific amino acid sequence present within them.
 RSS Feed
RSS Feed
